Yamada laboratory is interested in how mammalian cells touch, feel and work together to form multi-cellular tissues and organs. At the molecular level, cell-to-cell interactions are mediated by cell adhesion molecules and the cytoskeleton, and carefully regulated by accessory and signaling proteins. In particular, we are interested in how physical stimuli alter molecular interactions and regulation of cell adhesion, cell behavior, and ultimately cell fate. Since abnormal cell adhesion is common in diseases like cancer, understanding this unique force-dependent protein interactions should lead to novel treatments. If you are interested in our interdisciplinary research and would like to help, contact Dr. Soichiro Yamada for available positions. For more detailed descriptions of our research, please see our publications.
Our Research
Force-sensitive protein interactions

Physical force emerged as a key regulator of tissue homeostasis, and plays an important role in embryogenesis, tissue regeneration and disease progression. Currently, the details of protein interactions under elevated physical stress are largely missing, therefore, preventing fundamental, molecular understanding of mechano-transduction. This is in part due to the isolation of protein complexes under force-bearing condition is impossible using traditional solution biochemistry. Instead, our innovative biochemical analysis is based on in situ proximal biotin labeling using a promiscuous biotin ligase with a cell stretch device that promotes the formation of force-sensitive complexes.
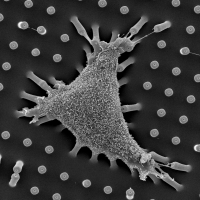
Using this approach, we identified a unique force-sensitive protein interaction surrounding α-catenin [Ueda el al.] and zyxin [Cheah et al.]. Currently, we are investigating other potential force-sensing machineries (e.g., cten) to uncover complete sets of force-dependent protein interactome, thereby resolving how force alters signaling cascade and cell phenotype. Based on our innovative strategy, we are in an ideal position to uncover the composition of force-sensitive complexes, a critical first step for understanding the molecular basis of mechano-transduction.
Self-contact induced membrane fusion
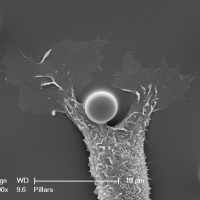
Biological membranes are frequently remodeled via membrane fusion and fission, a crucial requirement during tissue development and homeostasis. While intra-cellular membrane fusion is ubiquitous and well-studied, fusion between plasma membranes is rare and thought to be reserved for specialized cells. Using innovative micro-fabricated substrates, we demonstrated that mammalian epithelial and endothelial cells are highly fusogenic even in the absence of fusion-inducing factors [Sumida and Yamada and Sumida and Yamada]. However, this plasma membrane fusion is strictly limited to self-contacts, i.e., two extending and adhering membrane protrusions from a single cell, and not between two membrane protrusions from neighboring cells.
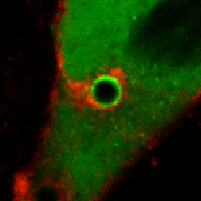
Our observations challenge an existing paradigm that the fusogenic capacity required for cell-to-cell fusion is reserved for specialized cells. Rather, fusion between plasma membranes is frequent, but limited to self-contacts and tightly regulated to avoid cell-to-cell fusion. Our intriguing observation suggests that epithelial and endothelial cells have a remarkably efficient molecular complex to fuse plasma membranes, and are capable of discriminating self- from neighboring cell-cell contacts. The obvious consequence of defective self-awareness is unwanted cell-to-cell fusion, a phenomenon observed in cancer cells and macrophages in tuberculosis. We are in process of identifying key molecules required for this unique membrane fusion.
